Abstract
Oceans are changing and these changes are affecting the animals that live there. Animals respond differently to changes in water temperature, food availability, and contaminants. Those responses can be seen in their genes. A technique called transcriptomics allows scientists to see the response of an animal’s genes to its environment. We used transcriptomics to compare two populations of Pacific razor clams in Alaska (United States): one that has lots of clams and one that used to have lots but does not anymore. We were surprised when we did not find any differences in their gene responses! So, we had to think about what else might be influencing the number of clams in these two populations. As we “dug” for answers, we found out that there are differences between the populations that do not influence their genes but may impact their numbers, such as being eaten by predators.
Genes Help Us Study Animals
Ecosystems around the world, including ocean ecosystems, are changing rapidly. How do we know if these changes are healthy or not? We cannot bring the ecosystems to a doctor for a checkup, but we can send scientists out to the ecosystems to measure the health of animal populations and individuals. By measuring repeatedly, scientists can tell how the ecosystems are doing. Comparing changes over time tells us what is changing and by how much. One way for scientists to measure ecosystems’ health is to measure the activity of the genes in the animals they study. Gene transcription is a cellular process that shows scientists how animals react to their environments, and studying gene transcription is one way to understand the health of the ecosystem.
Using Genes to Investigate a Razor Clam Population Crash
How do scientists use gene transcription to assess ecosystem health? Let us look at a real-world issue that has us scratching our heads: clams living on ocean shorelines. The clams we are studying are Pacific razor clams, a prized food for people who come to dig them and important prey for other animals that live along the coast, like sea otters. Historically, there have been two razor clam fisheries in Cook Inlet, Alaska (United States): one on the east side and one on the west side, about 63 km (or 40 miles) apart. About 10 years ago, the population of razor clams on the east side of Cook Inlet started to crash. The numbers became so low that scientists worried the population was in trouble. But the population of razor clams on the west side of Cook Inlet was doing fine. How could that be? These two populations of razor clams lived along the same body of water, in the same climate, in the same type of habitat, and close to each other (Figure 1). Scientists began to think about what could be different between the two populations.
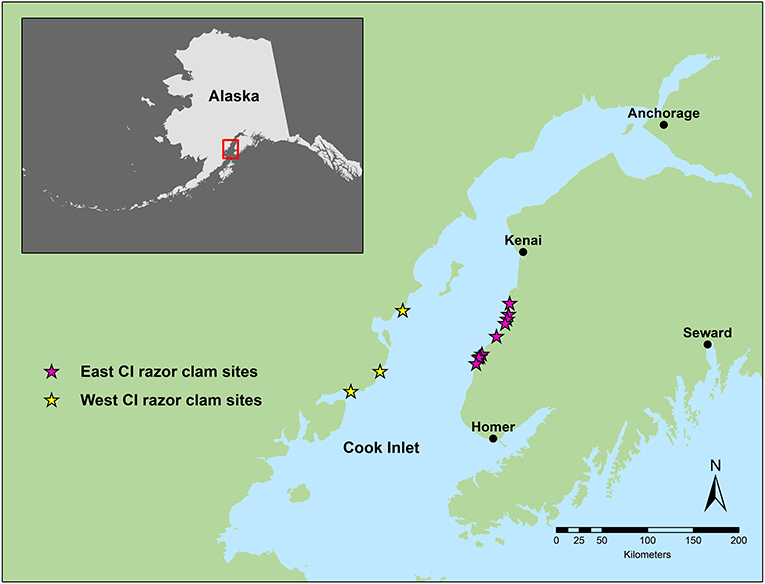
- Figure 1 - Map of Alaska and Cook Inlet (CI), where our study sites were located.
- The razor clam sites on the east side of Cook Inlet are shown with pink stars, while yellow stars show the west Cook Inlet razor clam sites.
Scientists wondered if there were differences between the two clam populations, like food availability, pollution, or increased ocean acidification. Some of these differences can be “seen” using a technology called transcriptomics. Transcriptomics measures how much a gene is turned on or turned off [1]. Genes have various functions in the bodies of every living thing, and genes can be turned on or off depending on whether or not they are needed at that time. Genes can be activated for many reasons, including certain types of stress. When genes are activated, DNA gets transcribed into a molecule called messenger RNA (mRNA). It is called messenger RNA because it carries DNA’s message. The mRNA gets turned into proteins that do important jobs within the organism. The amount of mRNA clams make depends on what is happening in their world. Let us say the razor clam encounters a contaminant, like fuel oil. The gene that makes contaminant-fighting proteins turns on, making more mRNA, which makes more proteins. Or, if a razor clam meets up with some bad bacteria, it will need to turn on its immune function genes for protection. By measuring the amount of mRNA that an organism makes from a specific gene, we can learn about problems razor clams may be facing (Figure 2).
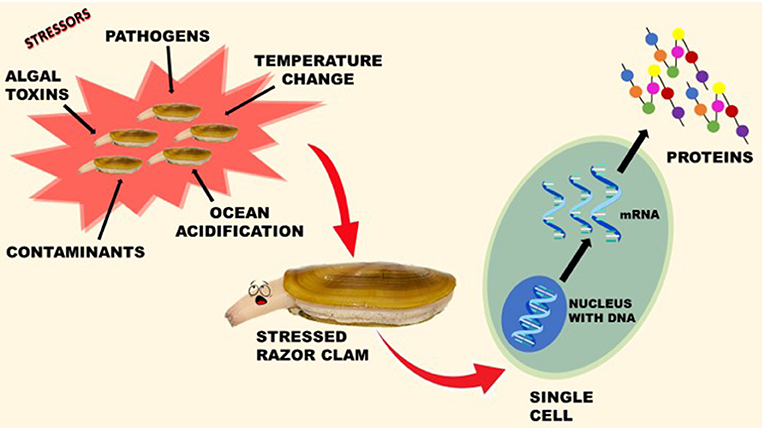
- Figure 2 - A razor clam can be stressed by many things.
- Stress activates genes, and DNA gets transcribed into mRNA. The mRNA gets turned into proteins that do important jobs within the organism. The amount of mRNA made depends on what is stressing (or not) the clam. Transcriptomics is the process of measuring the mRNA (figure credit: Min She Wenig).
How We Designed Our Study
Our goal was to monitor ecosystem health in Cook Inlet, where we were trying to figure out why there were different numbers of razor clams in two adjacent populations. We came up with a study plan to assess the health of the Cook Inlet ecosystem. We decided to collect 10 razor clams along nine sites on the east side and three sites along the west side of Cook Inlet, for a total of 120 clams per year. We wanted to look at their genes. We expected to find differences in the genes that were turned on and off between razor clams in east and west Cook Inlet. Those differences would tell us about the unique environmental pressures affecting each population.
First, we had to catch the clams. Using shovels, we dug holes on sandy beaches, hoping to find razor clams. We put the clams we found in a bucket and carried them back to the boat. Then we dissected them to get tissue samples for the gene analyses, which we did later, in the laboratory.
No Differences in Gene Transcription
We analyzed five genes that respond to stressors like pollution, ocean acidification, nutrition, and temperature. Somewhat surprisingly, we found no differences between the two sampling areas! The genes from razor clams living on beaches on both the west and east sides of Cook Inlet showed similar transcription patterns. So, what does that tell us?
There are two possible conclusions. First, maybe we did not test the right genes. There could be other genes that are affected by stressors that impact the health of the clams. So, we need to test more genes. In this study, we have only identified five razor clam genes, and this is a small number. In comparison, thousands of genes have been identified in humans! But human genes have been studied a great deal, and Pacific razor clam genes were not studied at all until we started our work. Second, maybe there are other differences between the two populations of clams that can affect their numbers, but do not make a difference to their genes.
It seems that something is affecting the east side razor clams, but not causing their genes to turn on or off. What could it be? Two ideas came to mind: habitat differences and predation. We needed to learn what researchers have discovered about habitat differences and predation between the two areas.
Could It Be a Change in Habitat?
We found that temperature and saltiness of the water are two things that might be different between the east and west sites. Algal blooms, which provide a burst of food for clams, are also probably different in timing and amounts of algae. However, if there were differences in food, temperature, or salinity that were big enough to affect the clams, we would expect to see gene transcription differences. Several of the genes we measured should turn on or off if the clams are stressed. Other habitat factors could involve big storms or changes to the sand in which the clams live. Storms and ice churn up beaches, remove sand, and can keep clams from reproducing or can even kill them. However, we would not see gene transcription changes in response to those kinds of events. In addition we did not find much information about storms and habitat loss over time in these two areas. More research is needed to answer this question.
Could It Be Predation?
We also wanted to look at predation—whether other organisms were eating the clams. Predation would not result in gene transcription changes. So, who could be eating the clams? We found that sea otters, a marine mammal that loves to eat clams and other marine invertebrates, live in Cook Inlet. Sea otters are a keystone predator [2], which means they have a large impact on their ecosystem through predation [3]. Sea otters have a very high metabolism, so they have to eat almost a third of their body weight every day. A 27 kg (60-pound) otter needs to eat almost 9 kg (20 pounds) of clams a day. That would be about the same as a 27-kg (60-pound) kid eating more than 20 pizzas every day! If sea otters were around, a lot of razor clams could be eaten, especially if there were not many other items on the otter menu in that area.
Scientists count sea otters in Alaska using airplanes. The data from Cook Inlet sea otter surveys were really surprising! We discovered that, in the last few years, the number of sea otters increased from <1,000 to over 3,000 on the east side. But on the west side, where the clams continue to thrive, there are still not very many sea otters.
While we did not start our study thinking about sea otters, we realized that the number of sea otters is very different between the east and west sides of Cook Inlet. Predation could be a big reason why there are far fewer razor clams on the east side of the inlet. We expect that sea otters and clam populations will eventually find a balance. To learn more about this balance, called carrying capacity, see this Frontiers for Young Minds article [4]. In the meantime, one of the next steps in our studies of razor clams will be to watch sea otters when they are eating. We want to understand how many sea otters there are and how many clams they are eating each year (Figure 3).

- Figure 3 - (A) Pacific razor clam after recently being dug.
- (B) Pacific razor clam samples bagged for analysis. (C) Scientists on a razor clam beach in Katmai. (D) Scientist sampling a razor clam: and (E) Sea otter eating a clam (photo credits: NPS Photo/J. Pfeiffenberger).
Conclusion
Our gene transcription results showed us that neither pathogens, contaminants, nutrients, or physiological stress seem to be causing the abundance differences between the east and west Cook Inlet razor clam populations. However, we now know that there are differences in the sea otter population sizes between the two areas and will conduct more studies to see what impact sea otters may be having on the clams. Our next steps for studying the clams will be to collect clams from the beaches, take them to a lab (alive) and conduct experiments to see if we can measure a response in certain genes by changing their food or the temperature of their water. Combined, these studies will allow us to have a better understanding of the impacts of environmental stressors, including predation, on the clam populations in Cook Inlet [5].
Funding
This work was supported by the National Park Service Foundation and the National Park Service. Alaska Department of Fish and Game provided support for sample collection at ECI. Lake Clark National Park and Preserve provided support for sample collection in WCI. Any use of trade, firm, or product names is for descriptive purposes only and does not imply endorsement by the U.S. Government. The data that support the findings of this study are available from the corresponding author upon reasonable request.
Glossary
Ecosystem: ↑ A community of organisms (animals, plants, and microbes) interacting with non-living things, like rocks and sand. Ecosystems can be as small as a puddle or as large as the ocean.
Gene Transcription: ↑ The process by which an RNA molecule copies a gene from the DNA so it can be used to build proteins.
Fishery: ↑ The harvest by humans of a marine species that is managed by a state or federal entity.
Ocean Acidification: ↑ Carbon dioxide (CO2) released in the atmosphere can be absorbed by the ocean. As the ocean absorbs CO2, the ocean waters are becoming more acidic (lower on the pH scale).
Transcriptomics: ↑ The scientific technique used for measuring gene transcription.
Dissect: ↑ To break something apart, or cut it up, to examine its parts and understand it better.
Algal Bloom: ↑ A rapid growth of microscopic algae in water. Algae are food for many marine animals such as clams.
Keystone Species: ↑ A species with a big impact on its ecosystem and that plays an important role in maintaining the ecosystem. Without it, an ecosystem might look very different.
Conflict of Interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
We thank Jim Bodkin for all his field and technical support. We also thank Jim Pfeiffenberger for his assistance with the photo collage (Figure 3) and the project in general and Min She Wenig for her artistry (Figure 2).
References
[1] ↑ Bourlat, S. J., Borja, A., Gilbert, J., Taylor, M. I., Davies, N., Weisberg, S. B., et al. 2013. Genomics in marine monitoring: new opportunities for assessing marine health status. Mar. Pollut. Bull. 74:19–31. doi: 10.1016/j.marpolbul.2013.05.042
[2] ↑ Estes, J. A., and Palmisano, J. F. 1974. Sea otters: their role in structuring nearshore communities. Science 185:1058–60. doi: 10.1126/science.185.4156.1058
[3] ↑ Paine, R. T. 1969. A note on trophic complexity and community stability. Am. Nat. 103:91–3. doi: 10.1086/282586
[4] ↑ Coletti, H., Hilderbrand, G., Bodkin, J., Ballachey, B., Erlenbach, J., Esslinger, G., et al. 2022. Where land and sea meet: brown bears and sea otters. Front. Young Minds 10:715993. doi: 10.3389/frym.2022.715993
[5] ↑ Coletti, H. A., Bowen, L., Ballachey, B. E., Wilson, T. L., Waters, S., Booz, M., et al. 2021. Gene expression profiles in two razor clam populations: discerning drivers of population status. Life 11, 1288. doi: 10.3390/life11121288
