Abstract
RNA, like its close cousin DNA, is used to store information in the cell. Unlike DNA, RNA is really good at folding up into interesting shapes, which also makes it good at lots of other important jobs. Some kinds of RNA, called riboswitches, can sense what is going on inside a cell. Each riboswitch fits a specific small molecule. When the riboswitch and small molecule interact, this changes what the cell does. For example, if the small molecule is harmful, the cell might start making a protein that will get rid of it. Recently, scientists discovered some riboswitches that look very similar to each other but recognize very different small molecules. We used a technique called X-ray crystallography to get pictures of these riboswitches, and we saw how changing just one piece of the riboswitch changed which small molecule it recognized. This shows us how RNA can gain new functions as an organism evolves.
A Day in the Life of a Cell
Being alive is hard work! All living things need to take in nutrients, grow, reproduce, and respond to an ever-changing environment. Many of the things we think about as being part of our environments are relatively large. We might notice a text message by hearing the phone beep, or an apple by seeing it hanging from a tree. But what about things that are too small to see or hear? For example, a bacterium might need to sense whether a toxic molecule is present so it can get rid of it, or it might need to know how much of a particular building block it has, so it can decide how to use the energy from its food. The molecules inside cells are extremely small—about one nanometer (one billionth of one meter) in size. Things this small cannot be seen with visible light, and they do not make any sounds. How can a creature notice something so small?
Somehow, organisms need to be able to “feel” the molecules they encounter. These molecules are crashing into each other many times each second, at speeds of hundreds of miles per hour. Molecules do not have much choice about what they crash into, but some pairs of molecules stick to each other better than others do. All organisms use this stickiness to their advantage, to help them get things done.
Anything You Can do, I Can do Better
Fifty years ago, scientists thought they understood the division of labor inside the cell. Proteins were the machines that did most of the work. Genes, the instructions to make proteins, were made of deoxyribonucleic acid (DNA). A close relative of DNA, called ribonucleic acid (RNA), carried those instructions to ribosomes, the cellular factories that make proteins. If this all seems too simple to be true, that is because it is. It is true that DNA is used primarily to store information, but RNA has many different jobs. This difference between DNA and RNA arises from a difference in structure (Figure 1). The two strands of DNA are almost always zipped up in one shape—the double helix. By contrast, RNA is single-stranded and can fold into a wide variety of shapes, depending on the sequence of its building blocks, which are called nucleotides. This makes it much like proteins, which also fold into unique shapes dictated by their amino acid sequences.
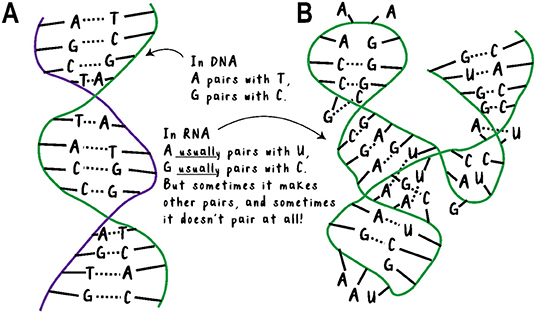
- Figure 1 - (A) DNA is a long chain of the nucleotides A, C, G, and T.
- It is almost always double-stranded and coiled up. Each nucleotide has a partner on the opposite strand to pair with, so there is not much room for variety in its shape. (B) RNA is a long chain of the nucleotides A, C, G, and U. It is usually found as a single strand, so the nucleotides do not have set partners on another strand. This leaves much more room for variety in both the structure and function of RNA. For example, the RNA in the picture is an enzyme, which can speed up a chemical reaction (Adapted from Scott et al. [1]).
RNA’s most familiar role is as a messenger. Messenger RNA (mRNA) is a temporary copy of a section of DNA. But other types of RNA can control how much protein is made, modify other molecules, and help the ends of chromosomes stay healthy. Even the ribosome, which manufactures all proteins, is made mostly of RNA! Some sequences of RNA fold up into a shape that allows them to stick to (or “bind”) certain small molecules. When a piece of RNA can bind to a specific small molecule, we call that RNA an aptamer.
Riboswitches: Aptamers With a Purpose
Who says an RNA must have just one function? Sometimes there is an aptamer in the front of an mRNA molecule. When the correct molecule binds to the aptamer, this can change whether the mRNA makes protein or not. These are called riboswitches (because they are on/off “switches” made from ribonucleic acid) and they work a lot like railroad switches (Figure 2). If there is a red signal, then the train cannot enter the main track, but if there is a green signal, then the train can start moving. If a riboswitch is “off,” it can prevent the ribosome from getting started making the protein. When the right small molecule binds to the aptamer, it changes the shape of the RNA so that the switch is “on” and the ribosome can start making protein.
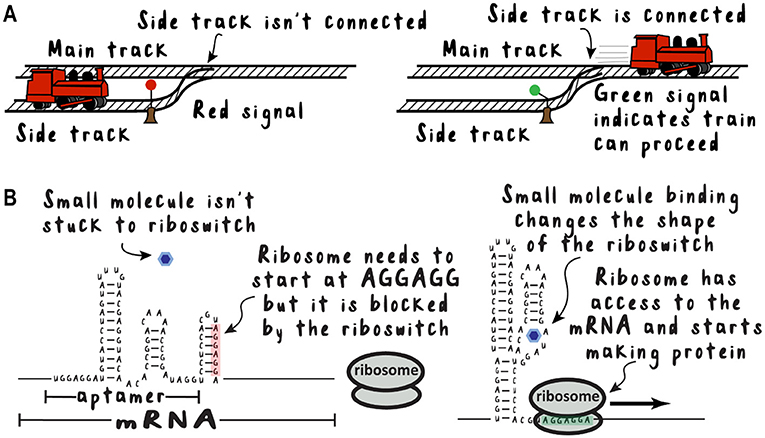
- Figure 2 - Riboswitches work like train switches.
- (A) A railroad switch responds to a signal by joining the side track to the main track, allowing the train to move to its destination. (B) A riboswitch responds to a small molecule by changing shape, allowing the ribosome to bind and start making protein.
Many microorganisms use riboswitches to sense various molecules within their cells and turn genes related to those molecules on or off. There are riboswitches that sense amino acids, nucleotides, vitamins, signaling molecules, minerals, and even fluoride. For a riboswitch to work properly, it needs to bind to just one kind of molecule but quickly let go of anything else it bumps into. This means the shape of the riboswitch must match the shape of the molecule it recognizes. How do riboswitches get their distinct shapes?
Family Differences
Like living organisms, RNA molecules can evolve from a common ancestor and become more diverse over evolutionary time. Scientists recently discovered a family of riboswitches that all have almost the same sequence and almost the same structure, yet they do different things. Some riboswitches in this family were found to bind guanidine, while other family members could not [2]. Guanidine is a small, toxic molecule with a positive charge. Those family members that cannot bind guanidine have almost the same sequence of nucleotides as those that can—but not exactly the same. Scientists were surprised to find that some of the non-guanidine-binding riboswitches can instead bind a molecule called phosphoribosyl pyrophosphate (PRPP) [3], while others can bind guanosine tetraphosphate (ppGpp) [4]. These two molecules perform different functions in the cell, they are both bigger than guanidine, and they have a negative charge instead of guanidine’s positive charge. ppGpp is a bit like PRPP with a G base from RNA or DNA added to it, but they are very similar otherwise. This fits with the fact that their abbreviations are also similar, which can be confusing (Figure 3A). Aptamers that bind PRPP do not bind ppGpp, and vice versa. This is like a lock that can tell your key from your neighbor’s by looking at the difference in just one pin. How does this work on the molecular level?
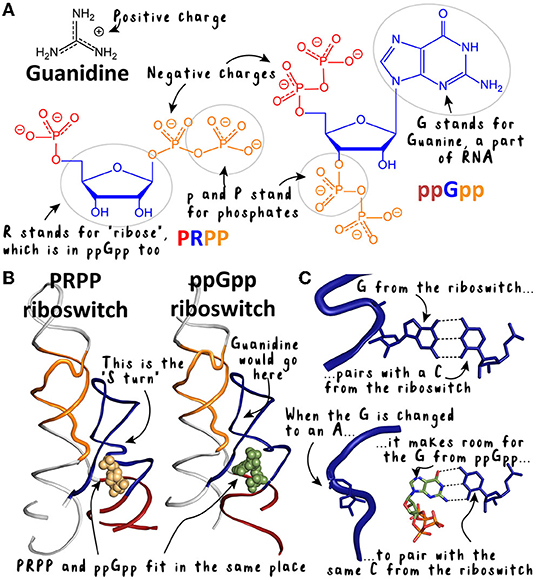
- Figure 3 - Structures of the PRPP and ppGpp riboswitches and the diverse molecules they bind.
- (A) Guanidine is small and positively charged, while PRPP and ppGpp are larger and negatively charged. The parts of PRPP and ppGpp are color coded to match their names. ppGpp is similar to PRPP with the “G” added on. (B) The PRPP and ppGpp riboswitches are like twins—they look almost identical. (C) If we zoom in to one place—the “S-turn”—we can see a difference in the ppGpp riboswitch that makes room for the G of ppGpp.
A Picture is Worth 1,000 Experiments
In the early days of biochemistry, it was not possible to see what proteins or nucleic acids looked like. Instead, scientists made careful changes to experimental systems and inferred what was going on at the molecular level based on their results. That approach is still important today, but we also have a rapidly improving ability to take pictures of molecules. In a study from our lab, we used X-rays to take pictures of an aptamer that binds PRPP [5]. This technique is called X-ray crystallography and it allows us to see the fine details of the RNA’s structure. We found that the PRPP-binding aptamer is bigger in the place where its binds its ligand, which is bigger than the guanidine ligand. This aptamer also uses a different area of its molecule to recognize guanidine [6]. Where guanidine binds in the guanidine-binding aptamer, the PRPP-binding aptamer has a metal ion (Figure 3B). This makes sense because the metal ion is also small and positively charged, like guanidine. Although this picture told us a lot about how the PRPP-binding aptamer interacts with its binding partner, we really wanted to know what keeps this aptamer from binding to ppGpp.
A Small Step in Rna Sequence, a Giant Leap in Specificity
The nucleotide sequences of the PRPP- and ppGpp-binding aptamers are extremely similar. In fact, we noticed that there was only one consistent difference. In one place—the 96th nucleotide of the PRPP-binding aptamer—there is always a guanine (G). In that same place, the ppGpp-binding aptamer almost never has a G. And ppGpp has an overall structure much like that of PRPP, except that it has a guanine base…could there be a link? We changed the 96th position in the PRPP-binding aptamer to an A, and kept all the other nucleotides the same. We then used X-ray crystallography to take pictures of this “mutant” aptamer to see how it works (Figures 3B,C) [5].
We saw that we could change the PRPP-binding aptamer into a ppGpp aptamer by changing position 96 from a G to an A. In the PRPP-binding aptamer, nucleotide 96 reaches across the structure of the molecule to base-pair with a C. When the aptamer is mutated to have an A at position 96, that A moves out of the way. This leaves the C free to pair with another G—in this case, the one from the ppGpp molecule (Figure 3C). Making this small change to the aptamer makes room for it to bind the larger ppGpp molecule. Guanidine is really different from both PRPP and ppGpp, so it takes a lot more changes to make an RNA that recognizes it [7].
Evolution in Miniature
When the scientist Charles Darwin visited the Galapagos islands, he found that there were many species of birds there that differed mostly in the size and shape of their beaks. The different species came about as descendants of the same ancestor. Some had beaks with slightly different shapes, making them slightly better at eating different kinds of food. If they were better at eating the food they found in their environment, then they were more likely to thrive and have more babies. In the same way, we think this family of riboswitches may have all come from the same ancestral RNA. Some of these ancestral RNAs may have ended up attached to the front of a gene, by chance. If an RNA adapted to respond to a molecule that made sense for that gene, then the bacterium would be more likely to survive and reproduce. So, the same way that Darwin’s finches look mostly similar but have different beaks, these RNAs also look the same but have differences in the size and shape of their binding pockets. This shows us that, just as organisms multiply and change in new environments, genetic elements within organisms do the same thing. This process has led to the existence of hundreds of riboswitches in bacteria. Each riboswitch can “see”—and respond to—something different, to help the bacteria survive.
Glossary
Ribonucleic Acid (RNA): ↑ A chain of nucleotides that includes the bases adenine, cytosine, guanine, and uracil.
Nucleotide: ↑ A building block of a nucleic acid. The most important part is the base, which determines what “letter” the nucleotide is.
Aptamer: ↑ A sequence of nucleic acid or protein that can bind to a small molecule.
Riboswitch: ↑ An RNA aptamer that changes the amount of protein made in response to binding a small molecule.
Guanidine: ↑ A small molecule that is toxic to cells. When there is too much in the cell, its proteins start to lose their proper shape.
Phosphoribosyl Pyrophosphate (PRPP): ↑ An important building block for how cells make nucleotides. It looks like a nucleotide with negative charges where the base should be.
Guanosine Tetraphosphate (ppGpp): ↑ An alarm signal for bacteria. It looks like a G nucleotide with more negative charges.
X-ray Crystallography: ↑ A method for figuring out the shape of nucleic acids, proteins, and small molecules by taking “pictures” of them with X-rays.
Conflict of Interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
We thank Caroline Reiss and Scott Strobel, the other authors of the original research paper, as well as all the members of the Strobel lab for their discussions and ideas. We also acknowledge support from the National Institutes of Health (Grant number GM022778 for the original research and GM136969 for the writing of this article).
Original Source Article
↑ Knappenberger, A. J., Reiss, C. W., and Strobel, S. A. 2018. Structures of two aptamers with differing ligand specificity reveal ruggedness in the functional landscape of RNA. eLife 7:e36381. doi: 10.7554/eLife.36381
References
[1] ↑ Scott, W. G., Finch, J. T., and Klug, A. 1995. The crystal structure of an all-RNA hammerhead ribozyme: a proposed mechanism for RNA catalytic cleavage. Cell 81:991–1002. doi: 10.1016/s0092-8674(05)80004-2
[2] ↑ Nelson, J. W., Atilho, R. M., Sherlock, M. E., Stockbridge, R. B., and Breaker, R. R. 2017. Metabolism of free guanidine in bacteria is regulated by a widespread riboswitch class. Mol. Cell 65:220–30. doi: 10.1016/j.molcel.2016.11.019
[3] ↑ Sherlock, M. E., Sudarsan, S., Stav, S., and Breaker, R. R. 2018. Tandem riboswitches form a natural Boolean logic gate to control purine metabolism in bacteria. eLife 7:e33908. doi: 10.7554/eLife.33908
[4] ↑ Sherlock, M. E., Sudarsan, S., and Breaker, R. R. 2018. Riboswitches for the alarmone ppGpp expand the collection of RNA-based signaling systems. Proc. Natl. Aacd. Sci. U. S. A. 115:6052–7. doi: 10.1073/pnas.1720406115
[5] ↑ Knappenberger, A. J., Reiss, C. W., and Strobel, S. A. 2018. Structures of two aptamers with differing ligand specificity reveal ruggedness in the functional landscape of RNA. eLife 7:e36381. doi: 10.7554/eLife.36381
[6] ↑ Reiss, C. W., Xiong, Y., and Strobel, S. A. 2017. Structural basis for ligand binding to the guanidine-I riboswitch. Structure 25:195–202. doi: 10.1016/j.str.2016.11.020
[7] ↑ Knappenberger, A. J., Reiss, C. W., and Strobel, S. A. 2020. A modular RNA domain that confers differential ligand specificity. Biochemistry 59:1361–6. doi: 10.1021/acs.biochem.0c00117
