Abstract
Every living thing needs food, also known as nutrients. In nature, living things are connected through food webs. Food webs show us how animals and plants are related through feeding relationships, like birds eating fish. But how do nutrients get returned back into the food web? The answer is bacteria! Bacteria are impossible to see with the naked eye. But, they are found almost everywhere in the world and are very important for recycling nutrients within habitats, like a river. Scientists use the DNA of bacteria to figure out which bacterial species are present and how these different bacterial species recycle nutrients in a river. Knowing which species are present helps scientists to use bacteria to monitor changes in river habitats. These changes might include the amount of food that is available for larger animals. Bacteria are considered to be the base, or foundation, of the food web, and paying attention to early changes in bacteria may allow us to prevent more extreme changes from causing problems later on for larger creatures.
Have You Ever Wondered Who Lives in the River?
Of course, we can imagine large creatures like fish swimming around in a river, but have you ever thought about the microscopic organisms that also live there, like bacteria? And what are the bacteria doing in the river, anyway?
Bacteria are small, single-celled organisms that are found on every continent and in almost every habitat, from rivers and oceans to soils and mountains. Bacteria even live on and inside other animals, like us humans. Most bacteria are smaller than the width of a human hair, and we use microscopes to help us to see their details. Once we can see them, it is obvious that there are many shapes and sizes (within a microscopic range) for bacteria, even in a single habitat like a river. Some bacteria are round (known as “cocci”), some are shaped like sticks (known as “rods”), and some are spiral-shaped (known as “spirochetes”). You can see examples of these bacterial shapes in Figure 1. Each species of bacteria is unique, just like humans are unique from other animals. Some bacteria may look very similar under a microscope, but are actually different species.
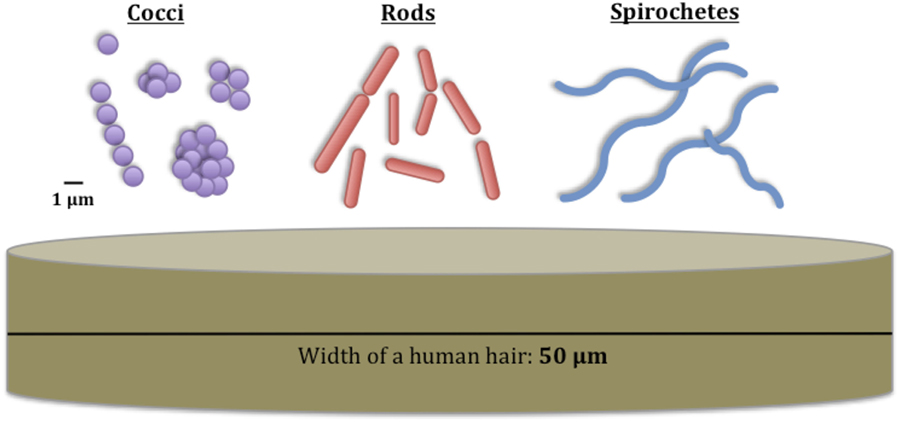
- Figure 1 - These are the different shapes of bacteria and their sizes compared with the width of a human hair.
- The unit “μm” is pronounced “micrometer” and is a very small measurement of length, 1/10,000 of a centimeter.
Why Does Anyone Care about Bacteria?
For many people, the word “bacteria” brings to mind images of germs and disease. But many bacterial species, including most of those in the river habitat, are harmless to humans. Bacteria are very important because they contribute to nutrient (food) cycles in many habitats. Because they are present in every habitat, bacteria are helpful indicators of a habitat’s health. In general, the health of a habitat depends on how well the organisms living there can survive. Just like humans need water, food and vitamins to grow, all other organisms need these same things. And bacteria have important roles in transforming the complex chemicals from organic matter into edible forms for other plants and animals. Organic matter is made of decomposed organisms like dead plant and animal tissues. Organic matter is very important because it is a food and energy source for living things in a river or any other habitat. When bacteria break down organic matter in the river, they help to fuel the aquatic food web.
Aquatic food webs are helpful in understanding the ways that producers and consumers of food are connected; you can see an example of a food web in Figure 2. In a river, birds eat fish and fish eat insects. Those insects eat plants and other smaller insects. And those smaller insects eat plants and bacteria. At the base of the food web, bacteria recycle the organic matter that is flushed from soil or dead plants and animals and put it back into the food web. Bacteria are very important to the food web, because they transform organic matter that other organisms cannot eat into edible food. Some organisms may also eat bacteria directly. In these ways, bacteria help to make up the base of the aquatic food web. You can imagine that, without bacteria and other decomposers, the organic matter from the food web would build up. Eventually, organisms higher on the food chain would have very little food available to them.
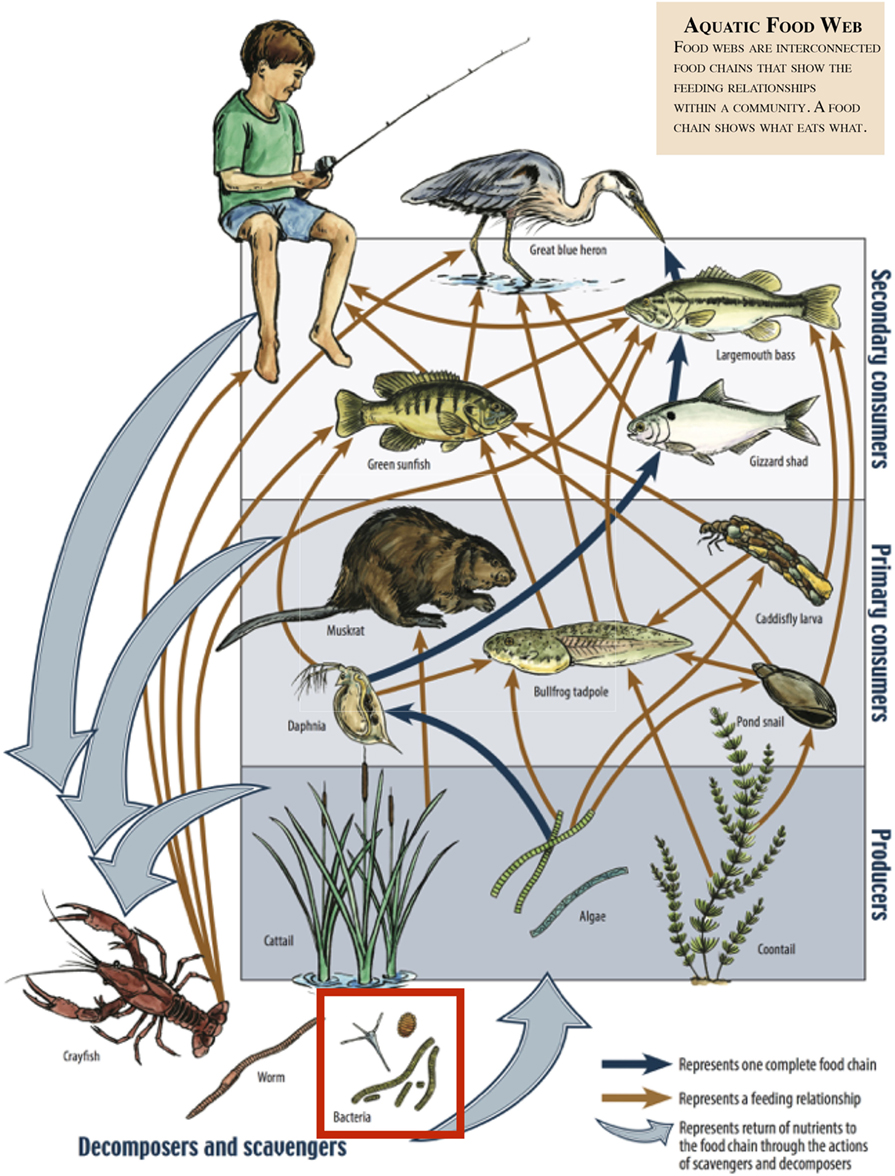
- Figure 2 - This is an example of an aquatic food web.
- Bacteria can be seen in the red box at the bottom. Bacteria (and other decomposers, like worms) decompose and recycle nutrients back to the habitat, which is shown by the light blue arrows. Without bacteria, the rest of the food web would starve, because there would not be enough nutrients for the animals higher up in the food web. The dark orange arrows show how some animals consume others in the food web. For example, lobsters may be eaten by humans. The dark blue arrows represent one complete food chain, beginning with the consumption of algae by the “water flea” Daphnia, which is consumed by a small fish, which is consumed by a larger fish, which is at the end consumed by the Great blue heron. [Image (amended) was provided courtesy of the Missouri Department of Conservation.]
Because they are so important for nutrient recycling at the base of the food web, bacteria can be helpful for indicating habitat health. Scientists have found that these bacterial communities react quickly to changes within a habitat, such as water temperature changes or changing amounts of organic matter in the river [1]. These changes may have important consequences for the larger animals, like birds and fish. Certain changes may help some species to survive, but they may also make life harder for other species. But, by the time the changes affect larger animals, the habitat has probably already been changed to a dangerous level. In a river, these changes might be warmer water temperatures or increased amount of organic matter. Unusual changes to water temperature can be harmful for some bacteria. Changes in their living conditions may create problems for those bacteria that provide food and nutrients to other animals in the habitat. If there is not enough food for the larger animals in the river, there can be other problems that follow. These problems could be predicted ahead of time, using the health of the bacteria as measure of habitat health. The types and numbers of bacteria can be monitored to help warn scientists of small changes to the habitat that may, in the future, have a larger impact on the habitat.
Do Scientists Know How Many Bacterial Species are There in a River?
Scientists have found that one teaspoon of river water can have millions of bacteria [2]! But it is difficult to count the exact number of bacteria in any habitat. Scientists can estimate the different species. Scientists are also able to describe these organisms based on how they survive, or their lifestyles.
Each bacterial species is unique and different species have different jobs for recycling organic matter in a river habitat. The bacterial species can be unique in the way they look, swim, or where in the river they live. For example, some bacteria attach themselves to particles in the river water. These particles are generally small pieces of clay, sand, or dead plant material and they make rivers murky or dirty looking. But the particles can be a great place for bacteria to hang onto, as well as a food source. The bacteria on particles are called “particle-attached” bacteria. Other types of bacteria drift or swim on their own in the river water. They are known as “free-living” bacteria. Our team was curious to find out if bacteria with these two different life styles, particle-attached and free-living, have different jobs or roles in nutrient recycling in the river. We examined different species of bacteria living in the Columbia River and estuary (the estuary is the lower part of the river, where the river water and the ocean tides meet and mix together) in the Pacific Northwest of the United States and in the nearby Pacific Ocean.
Did the Scientists Find Differences Between the Roles of Particle-Attached and Free-Living Bacteria in the River?
Yes, particle-attached and free-living bacteria have different lifestyles and they contribute to the aquatic food web in different ways. Higher numbers of particle-attached bacteria are found in murky, dirty-looking water [3]. This makes sense, because murky water has more particles in it and more places for bacteria to attach. So, water with particles may seem gross and dirty, but it is home to bacteria with important jobs. Our team also found that, in clearer water without particles, free-living bacteria were more widespread. So, the different aquatic habitats select for different lifestyles of bacteria, and nutrient recycling occurs in both places!
To make sense of these different lifestyles, we can compare the different river lifestyles of bacteria to people on ships or boats. Just like on a ship, bacteria can be situated closely together on the particle surface. On a particle, different kinds of bacteria can be near each other and perform different jobs. Like in the ship’s kitchen, one chef might be cutting up carrots while another chef is adding the finishing spices to a stew. Just like these chefs, some bacterial species seek out organic matter and transform it into something that another organism can eat. We can imagine the bacteria doing similar activities on a particle. Some bacteria are picky and eat only very specific nutrients. Other bacteria cannot break down the more complex food. Instead, they have to find simple, ready-to-eat food from the river water. Other bacteria are not so fussy and will consume anything they can get ahold of. After consuming organic matter, the bacteria (and maybe even the particle where they live!) are often eaten by larger, but still microscopic, organisms.
We know that bacteria are important to a habitat, but they are hard to see with our naked eyes. To understand what the bacteria can do, our team studied the genes of these bacteria. We found that particle-attached bacteria also appear to have a larger number of genes, or instructions for living, compared with free-living bacteria. We looked at the different types of bacterial genes to understand better how the bacteria live their lives in the river. For example, particle-attached bacteria have more genes used to break down and consume small, photosynthetic organisms (known as “phytoplankton”) than the free-living bacteria have [4]. This means there are more instructions telling particle-attached bacteria how to recycle dead phytoplankton. This makes sense, since we find more particle-attached bacteria in places with higher amounts of organic matter and more particles, both of which can come from dead phytoplankton. However, particle-attached bacteria are not as widespread in clear water as are free-living bacteria. So, it appears that particle-attached bacteria are important for recycling organic matter in murky river habitats. It also appears that free-living bacteria are important for nutrient cycling in clear-water river habitats. With this information, we know a bit more about how and where bacteria with different lifestyles contribute to aquatic food webs in rivers. So, if in the future we see some unexpected changes in the bacterial species living in murky or clear river waters, this can tell us that the river habitat is changing, even before birds and fish higher in the food web are affected by the changes.
But How Can We Tell if the Bacterial Species in a Habitat are Changing?
When we look through a microscope at bacteria, we can only see what bacteria look like. Seeing shapes of bacteria may be helpful to tell one type apart from another, but there are many different types of bacteria that play different roles in the environment that look very similar to each other under a microscope. In fact, looking at their shapes tells us very little about what species they are or how they contribute to the aquatic food web. To find out exactly which species a bacterium is or what jobs it performs in the river, we examine its DNA. DNA is a code that is a kind of fingerprint for every species. DNA is made up of four chemical building blocks, referred to as bases. DNA bases are abbreviated as the letters A, T, G, and C. DNA makes up genes, or the instructions for each organism’s structure and lifestyle. We can read the DNA letters as a code to understand bacterial genes. The genes give us information about which species a bacterium is and what it can do in the river.
Our team collected the DNA from bacteria attached to particles and from free-living bacteria. We used two filters with very small holes that were stacked together to separate particles and free-living bacteria from the river water. Particles can easily be filtered out of water; you can think of water purifiers that are used to clean up water. But in this case, we are interested in what is caught in the filter, not in the clean water. Remember, bacteria are small, but water molecules are even smaller. So the particles with the particle-attached bacteria are collected with the first filter with holes of 3 μm in diameter (a μm is 1/10,000 of a centimeter) and the free-living bacteria are collected in the second filter, which has even smaller holes (0.2 μm). We used these filters to capture the bacteria for extracting and purifying their DNA. Extraction and purification of DNA involves breaking open the bacterial cell walls and adding a series of chemicals to separate the DNA from the bacterial cell wall materials and other cell components, like proteins. It is important to remember that the purified DNA is actually a mixture of all the DNA from all of the different species of bacteria we collected in our water sample.
Once we had our mixture of pure bacterial DNA, our team used a technique called metagenomics, or the study of all the DNA in water samples, to find out which bacteria were there and what they were doing in the river. The DNA was put through a large sequencing machine that reads and records the DNA codes of each sample. The DNA codes were compared to known lists of other bacterial species’ DNA. There are sections of DNA that have not changed very much over a long period of time through evolution. Those are called conserved DNA regions. Letter changes (DNA base changes) in those conserved DNA regions help us to tell species apart. These DNA codes help us to understand which species of bacteria are present in the water samples, kind of like an attendance sheet for bacteria.
DNA helps us identify which species are present in a water sample, but it also helps us understand what jobs bacteria are able to do in their habitat. The bacterial DNA codes can be compared to instruction manuals that help us to understand what the bacteria are capable of doing [5]. These instruction manuals tell each species which jobs they can perform, such as how to find nutrients and which kinds they can consume. Consuming different kinds of nutrients requires different instructions. For example, bacteria that make food from sunlight (called “photosynthetic” bacteria) have different instructions coded into their DNA compared with other bacteria that break down complex organic matter, like wood or plant material. Knowing about the different bacterial jobs required to recycle nutrients in a habitat is helping scientists to understand how important bacteria are within aquatic food webs.
Our team wants to understand how aquatic bacteria react to changes in the environment, especially before those changes impact larger animals in the aquatic food web, like birds and fish. We were one of the first scientific groups to use metagenomics to study river bacteria using filters with different sized holes. Our team’s discoveries help scientists to understand more about the kinds of bacterial species and different jobs of free-living and particle-attached bacteria in aquatic habitats. Knowing more about the bacteria at the base of the food web helps scientists to identify and possibly prevent the negative impacts of environmental changes in aquatic habitats. If studying the bacteria in the habitat can help us catch environmental changes early, hopefully those food web changes will be less likely to have long-term negative impacts on other organisms in aquatic habitats. Using the power of metagenomics, scientists can make better decisions about keeping river habitats healthy for human use, as well as for other important creatures like fish and birds.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Original Source Article
↑ Simon, H. M., Smith, M. W., and Herfort, L. 2014. Metagenomic insights into particles and their associated microbiota in a coastal margin ecosystem. Front. Microbiol. 5:466. doi:10.3389/fmicb.2014.00466
References
[1] ↑ Fortunato, C. S., Eiler, A., Herfort, L., Needoba, J. A., Peterson, T. D., and Crump, B. C. 2013. Determining indicator taxa across spatial and seasonal gradients in the Columbia River coastal margin. ISME J. 7, 1899. doi:10.1038/ismej.2013.79
[2] ↑ Maranger, R., and Bird, D. F. 1995. Viral abundance in aquatic systems: a comparison between marine and fresh waters. Mar. Ecol. Prog. Ser. 121, 217–226. doi:10.3354/meps121217
[3] ↑ Crump, B. C., Baross, J. A., and Simenstad, C. A. 1998. Dominance of particle-attached bacteria in the Columbia River estuary, USA. Aquatic Microb. Ecol 14, 7–18. doi:10.3354/ame014007
[4] ↑ Smith, M. W., Allen, L. Z., Allen, A. E., Herfort, L., and Simon, H. M. 2013. Contrasting genomic properties of free-living and particle-attached microbial assemblages within a coastal ecosystem. Front. Microbiol. 4:120. doi:10.3389/fmicb.2013.00120
[5] ↑ Markowitz, V. M., Ivanova, N. N., Szeto, E., Palaniappan, K., Chu, K., Dalevi, D., et al. 2007. IMG/M: a data management and analysis system for metagenomes. Nucleic Acids Res. 36(Suppl 1), D534–D538. doi:10.1093/nar/gkm869
