Abstract
When you hear the word microbes, what comes to your mind? Something much too small to see and that makes you fall ill? Just because some microbes cause diseases that does not mean they are all evil. For example, in the marine (ocean) environment, the vast majority of microbes are good ones. They are the “driving engines” of the ocean and are essential for the health of our whole planet. Unfortunately, most of the marine microbes and their interactions with the marine environment are poorly understood. So, it is important to get an idea of which microbes are helping us and how they are doing this. These data will provide scientists with the knowledge to fight against big global challenges, such as climate change and environmental pollution. Unfortunately, it is very hard to study marine microbes due to their microscopic size, huge diversity, and their big home – the ocean. Therefore, we would like to engage “citizen scientists” in this project to help us to sample marine microbes so that we can identify them.
Marine Microbes, the Driving Engines of the Ocean
When you hear the word microbes, what comes to your mind? Something that we cannot see with the naked eye? Invisible organisms that can invade our bodies and make us fall ill? Almost everyone has heard about bad things that microbes can do, such as cause dental cavities – the breakdown of teeth caused by bacteria. Pertussis, also known as whooping cough, is another well-known bacterial disease. The dislike that humans generally feel toward microbes is strong and in many cases understandable. However, this does not mean that all microbes are all evil. It may sound strange, but most of the microbes that exist are actually very helpful. Many of the “good” microbes can be found in the ocean (Figure 1), among other places. These marine microbes are microscopic single-celled organisms that include bacteria and archaea. Archaea are bacteria look-alikes, but with some special features such as the ability to live in extreme environments, for example, in extremely hot or extremely cold places. Moreover, marine microbes also include single-celled organisms, such as microscopic algae and even viruses. Marine microbes are tiny, but they exist in very large numbers and can be found all throughout the ocean, from the deepest parts up to the surface layers. It has been estimated that just one drop of seawater contains millions of microbes! In other words, there are more marine microbes in the oceans than there are people living on earth. In addition to being the unseen majority in the ocean, which is the world’s biggest ecosystem, marine microbes also form the basis of the marine food web, are responsible for the recycling of nutrients, and are involved in all types of important activities. For this reason, they are also called the “driving engines” of the ocean, meaning that they are essential to functioning of the ocean and the ocean cannot survive without them. Because they are a part of so many different processes, marine microbes have an important impact on our daily life and well-being, no matter where we live. One example of a function performed by marine microbes that influences humans is photosynthesis, a process you might have heard of in connection with plants. Like plants, some marine microbes, like Cyanobacteria, can use the light energy from the sun to convert carbon dioxide (CO2) and water into sugar. During this process, oxygen (O2) is produced and released into the environment. It has been estimated that about half of the world’s oxygen production takes place in the ocean, while the other half comes from terrestrial habitats. In other words, marine microbes are responsible for the oxygen in every second breath you take. Another example of the importance of microbes in the ocean is the ability of some microbes to break down oil. Some marine microbes feed on oil, which can be very useful for clean-up after an oil spill.
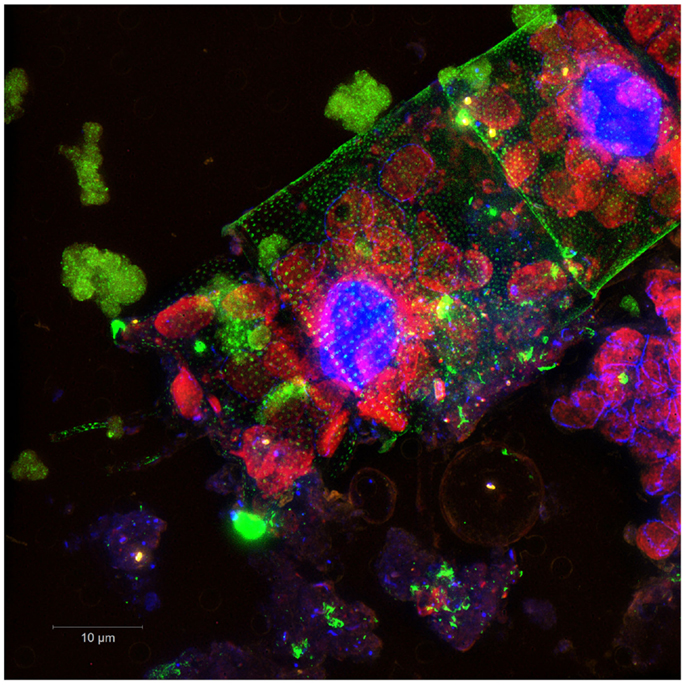
- Figure 1 - Image illustrating the microbial diversity in the ocean. It shows diverse microbial cells located on and around an algae. The microbes have been visualized using a colored dye. Image provided by Gomez-Perreira and Fuchs. (Picture courtesy of P. Gomez-Perreira and B. Fuchs, Max Planck Institute for Marine Microbiology, Bremen, Germany).
So, it is obvious from these examples that marine microbes are not just the driving engines of the ocean, they are also essential for the health of humans and the whole planet. The interaction between these microbes and the larger organisms on the planet is in a delicate balance, and could be threatened by factors, such as pollution and changing environmental conditions. In order to protect and maintain a healthy environment, we need to gain deeper knowledge about the kinds of microbes that exist, what are they doing, and how they interact with each other and with the environment. Unfortunately, collecting this information is not as easy as it sounds.
Why Do We Know so Little about Marine Microbes?
Until recently, scientist needed a pure culture of microbes to get an answer to simple questions, such as what they are like and what functions they are capable of. A pure culture means that a single microbe has to grow in the lab in the absence of any other species. Since conditions in the lab are very different from those in the oceans, it is extremely difficult to grow marine microbes. It has been estimated that only around 1–10% of marine microbes can be cultured in the laboratory [1]. We still do not know the exact reasons for this difficulty, but luckily, over the past few years, new techniques have been invented to allow the study of marine microbes without the need to grow them in pure culture in the lab first. One of these techniques is called Next Generation Sequencing (NGS).
How Does NGS Help Scientists to Understand Marine Microbes?
All the information about a microbe, or any kind of cell, exists in the cell’s DNA. DNA is the molecule that contains the genetic code of organisms and, thus, can be described as the “blueprint” of the cell, since it tells the cell what to do and when to do it. In can further be divided in small subsections called genes. There are thousands of genes in the DNA of an organism and each gene has a specific function. For example, the human DNA contains about 25,000–35,000 genes, but only a very few genes are responsible for the color of your eyes. The NGS technology allows scientists to “read” the DNA from a whole microbial community without the need for pure cultures of the microbes. This approach is called “Metagenome sequencing” and it provides the scientist with a list of the genes of all the microbes living in a particular area.
Some genes exist in all organisms on earth but have small differences between organisms. One example of this type of gene is the one for “ribosomal RNA” (rRNA). Because this gene is unique for each species, scientists can use it as a kind of fingerprint of a microbial species. You might have heard of fingerprint analysis being used as part of a criminal investigation. Law enforcement offices, such as the police, store all fingerprints in large databases for comparison with fingerprints taken at crime scenes. This allows the police to identify people who commit crimes. Scientists do the same with the gene fingerprint of microbes obtained through research. They use the rRNA gene to search in a specific database, such as the SILVA [2] database, for a positive match. With this approach, they can identify the microbes obtained in their samples and find an answer to the question “What kind of marine microbes are out there?”
But the DNA contains so much more than just a fingerprint. As mentioned above, it is the building plan of an organism and includes instructions for all of that organism’s features and functions. If scientists are able to “read” the data coded in the DNA, they can obtain a great deal of knowledge about that organism. Unfortunately, deciphering the DNA data obtained from marine microbes is like trying to understand an ancient language or to put together a puzzle. It takes a while, and the scientists might need to gather some additional “pieces,” but sooner or later they are able to solve it. Every day scientists discover a new piece of the puzzle and more and more information about marine microbes is revealed. Although NGS is still a fairly new method, this technology is clearly helping us to reach a new level of understanding of marine microbes from all parts of the ocean. However, the knowledge we have gained so far about marine microbes is just “a drop in the ocean” – there is so much more to learn! The Ocean Sampling Day (OSD) project [3] aims to greatly increase our knowledge of marine microbes by bringing people together to study the microbes in ocean ecosystems all around the world.
The Ocean Sampling Day
Researchers working on a project called Micro B3 (Marine Microbial Biodiversity, Bioinformatics, Biotechnology)1 organized an OSD [3] on June 21 in both 2014 and 2015 with the aim of studying marine microbes worldwide with the help of a large international group of scientists. June 21 is known as the solstice, meaning that it is the longest day of the year in the northern hemisphere and the shortest day of the year in the southern hemisphere. The solstice was chosen for microbe collection to see if day length influences the kind of microbes that are collected [4]. For the OSDs events in June 2014 and 2015, scientists around the world collected marine microbial samples. Each team filtered sea water through a specific filter called Sterivex? that contains very tiny holes that are smaller than microbes, so the microbes get stuck on the surface of the filter. This is how scientists capture microbes from the ocean or any aquatic system. Next, the scientists removed the microbes from the filter and used some chemicals to collect the DNA from all of the organisms obtained. Then, the DNA information was analyzed with the NGS technology.
The aim of OSD is to answer several questions: “what kind of microbes are out there?” (biodiversity analysis based on the DNA “fingerprint”); “what are those microbes doing?” (functional analysis based on metagenomics); and “how do those microbes interact with each other and the environment?” (Figure 2). Scientists are still analyzing the OSD data, so there are no reliable results to show yet. Hopefully, the data we obtain from OSD will help us to understand things such as how microbes differ between tropical and polar environments and how microbes can survive in contaminated environments, such as after oil spills.
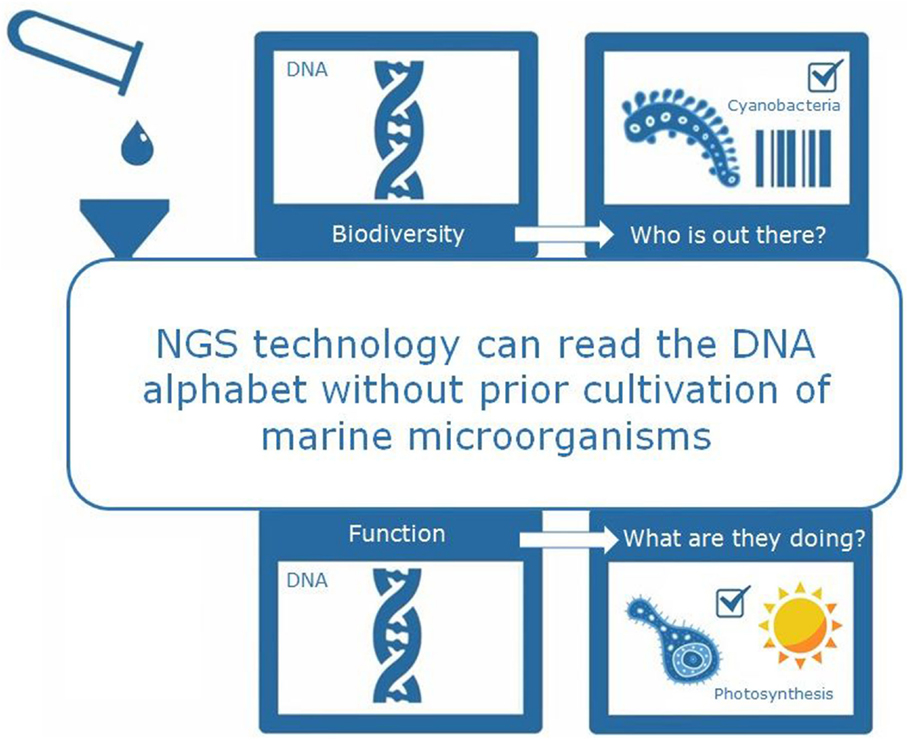
- Figure 2 - Next generation sequencing (NGS) technology is a powerful tool. It allows scientists to rapidly read the DNA from millions of microbes without prior culture in the lab. Microbial DNA is obtained from marine microbes in the ocean water samples and can be read using the NGS method. The DNA, as the “blueprint” of the microbial cell, contains all the information we need to answer questions such as “Which microbes are out there?” (Biodiversity) and “What are those microbes doing” (Function).
Humans have a very close and important relationship with the ocean. Not only does around half of the world’s population live within 200 km of the coastline [5], but the ocean is also important to the economy as a place for tourism, recreation, fishing, and other activities. All of these human actions affect and change coastal regions. As the majority of the OSD sampling sites are located in coastal areas (Figure 3), we want to study the ways in which human activities and lifestyles influence marine microbial communities. For example, we can study the way that various types of pollution (heavy metals, antibiotics, and fertilizers) affect the health of marine microbes.
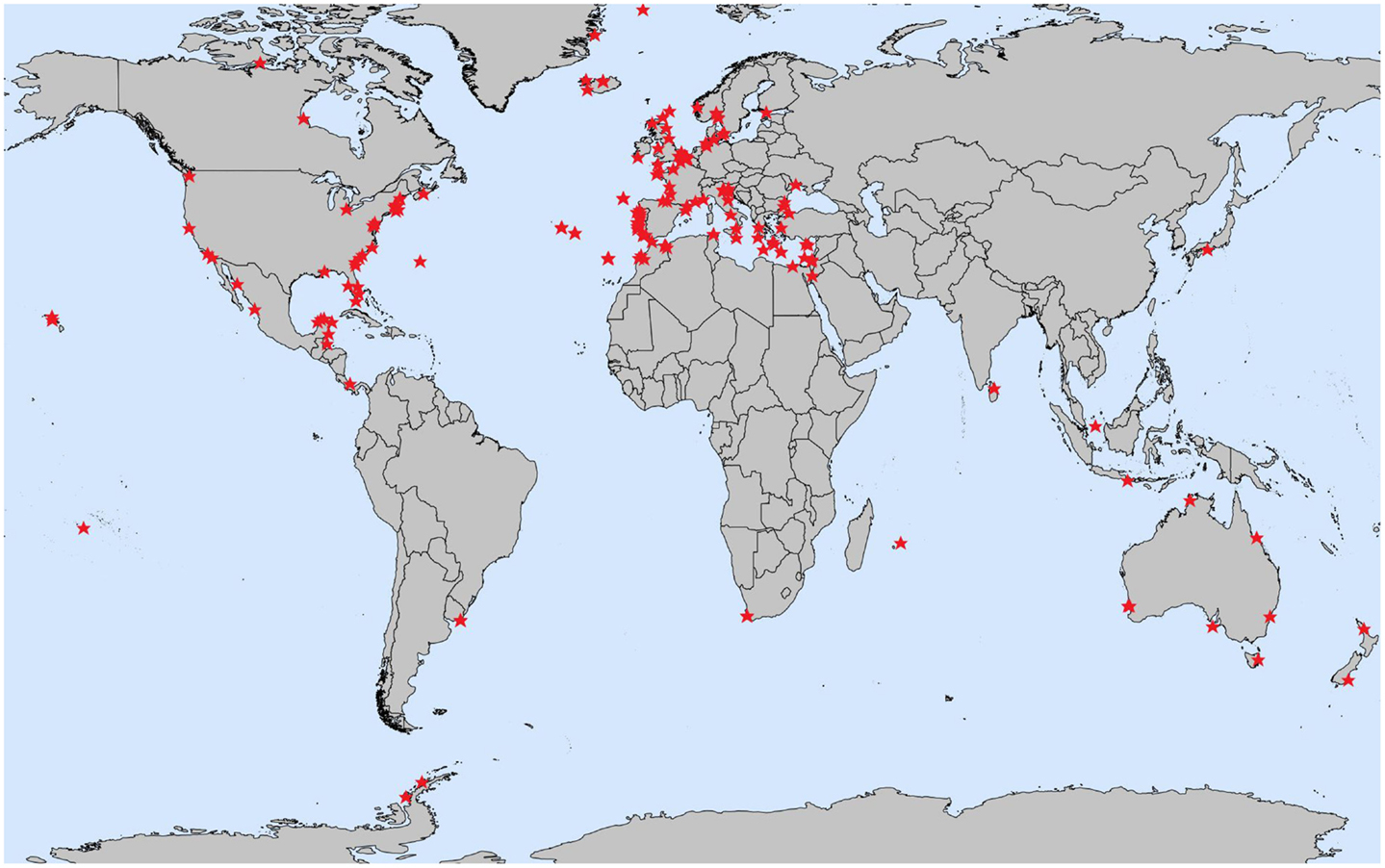
- Figure 3 - Map showing registered sites for OSD 2014. These sites are typically located in coastal regions. The red stars mark the locations at which scientists went out to collect seawater samples during Ocean Sampling Day.
Did you know that more than 70% of the earth surface is covered by the ocean [6]? As we move forward, we will continue expand the scope of OSD to explore more of the ocean. OSD 2014 and 2015 already involved more than 150 scientific teams from all continents, collecting samples from subtropical waters in Hawaii to polar environments, such as the Fram Strait in the Arctic Ocean. However, there are still many unexplored areas on the OSD map (Figure 3). To increase our efforts, we also invite “citizen scientists,” like you, to join our citizen science project called MyOSD. We developed a MyOSD Sampling Kit and an OSD Citizen Smartphone App, which allows citizen scientists to make a real contribution to science. The kit contains all the equipment anyone would need to collect marine microbes, as well as other material to measure additional important data, such as water temperature and salinity (saltiness) (Figure 4). Each kit comes with clear directions describing each step in detail. The concept is easy and straightforward, meaning that everyone can join MyOSD. Sampling kits are available for free through registration with MyOSD, and are distributed by MyOSD “hubs” or centers, to citizen scientists in their area or country. A list of all hubs can be found on the MyOSD homepage.2 After collecting samples, the citizen scientists need to return their samples to the MyOSD hub, which will send them all back to the OSD headquarters at the Max Planck Institute for Marine Microbiology3 in Bremen, Germany. Here, all the samples are processed and data analysis is coordinated. If there is no MyOSD hub in your area, you can still participate by measuring environmental parameters, such as water temperature and/or salinity. All data can be submitted via the OSD Citizen Smartphone App or the online form available on the website. Every data point counts! For example, if your data help to identify a unique temperature profile for a certain marine area, which might get the attention of scientists who will then design follow-up studies. In the end, all data will be freely available for everyone on the Internet, so that the OSD data can help generations of researchers and citizens. OSD 2014 and 2015 marked just the beginning of this project, and we hope to engage even more people for the upcoming event in June 2016!
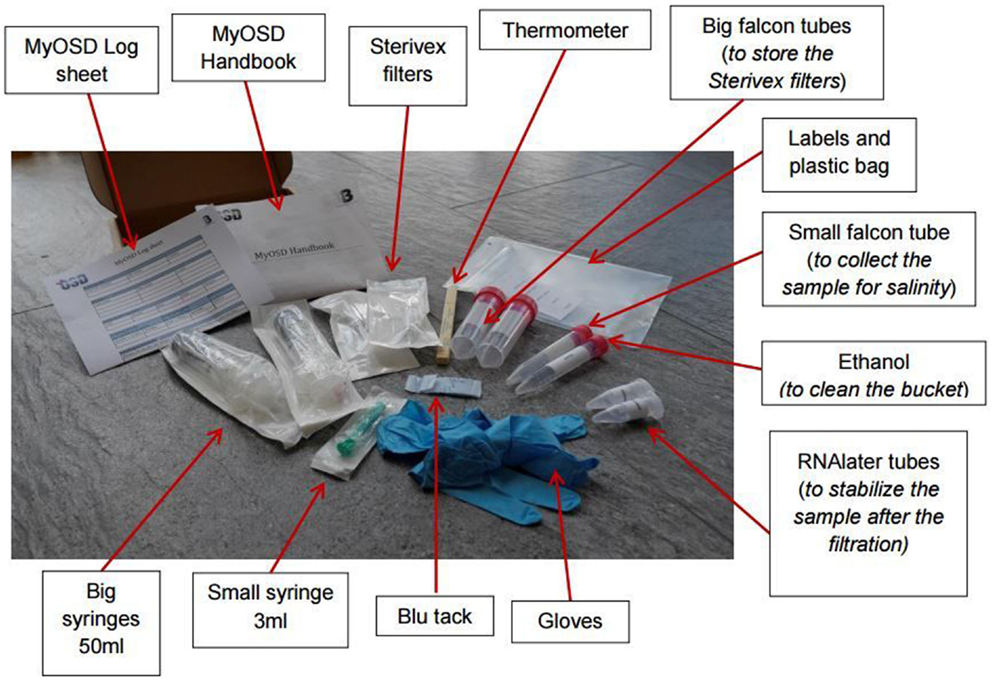

- Figure 4 - Contents of the MyOSD sampling kit. The materials provided in the kit include all of the necessary supplies a citizen scientist needs to collect samples of ocean water that can then be sent back to headquarters and analyzed by OSD.
Glossary
NEXT GENERATION SEQUENCING (NGS): ↑ This is a fairly new method for reading and recording the DNA. It allows scientists to read large amounts of the DNA letters with and without growing the organisms in a lab. Sometimes it is also called massive parallel high throughput sequencing.
DNA: ↑ Abbreviation for deoxyribonucleic acid. Can be described as the instruction manual or “blueprint” of every microbe or any other living cell. The information is stored as a sequence of four different molecules that are represented as letters (A, G, C, T).
GENE: ↑ Each DNA contains subsections called genes, which tells each cell what to do and when to do it. There are thousands of genes in the DNA of an organism and each gene has a specific function.
SEQUENCING: ↑ In genetics, sequencing means to read and record the sequence of letters (A, G, C, T) of the DNA of a cell.
rRNA GENE: ↑ This gene codes for the “ribosomal RNA” of a cell. The rRNA gene is very important for the cellular metabolism by playing a key role in the production of proteins. Every organism on earth has this gene in its DNA and it is unique for each species; thus, it can be used as a marker for the genetic fingerprint.
BIODIVERSITY: ↑ This term describes the variety of different types of life found on earth.
METAGENOMICS: ↑ This term describes the study of DNA recovered directly from environmental samples. The resulting data set contains the DNA of all microbes in the sample and not just a single species.
Footnotes
[1] ↑ www.microb3.eu
References
[1] ↑ Amann, R. I., Ludwig, W., and Schleifer, K. H. 1995. Phylogenetic identification and in situ detection of individual microbial cells without cultivation. Microbiol. Rev. 59, 143–169.
[1] ↑ Quast, C., Pruesse, E., Yilmaz, P., Gerken, J., Schweer, T., Yarza, P., et al. 2013. The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res. 41, D590–D596. doi: 10.1093/nar/gks1219
[3] ↑ Kopf, A., Bicak, M., Kottmann, R., Schnetzer, J., Kostadinov, I., Lehmann, K., et al. 2015. The ocean sampling day consortium. Gigascience 4, 27. doi: 10.1186/s13742-015-0066-5
[4] ↑ Gilbert, J. A., Somerfield, P., Temperton, B., Huse, S., Joint, I., and Field, D. 2010. Day-Length Is Central to Maintaining Consistent Seasonal Diversity in Marine Bacterioplankton. Nature Precedings. Available at: http://precedings.nature.com/documents/4406/version/1
[5] ↑ Creel, L. 2003. Ripple Effects: Population and Coastal Regions. Washington, D.C: Population Reference Bureau, Measure Communication.
[6] ↑ Costello, M. J., Cheung, A., and De Hauwere, N. 2010. Surface area and the seabed area, volume, depth, slope, and topographic variation for the world’s seas, oceans, and countries. Environ. Sci. Technol. 44, 8824–8828. doi: 10.1021/es1012752
